NESTML Izhikevich tutorial
Introduction
The aim of this exercise is to obtain familiarity with NESTML by completing a partial model of the Izhikevich neuron [1].
Prerequisites
You need to have a working NEST Simulator and NESTML installation, see Installing NESTML.
You need to be able to run NESTML to generate and build model code. NESTML can be used from the command line, and via a Python API. The latter will be used in this notebook. See Running NESTML.
[1]:
%matplotlib inline
import matplotlib.pyplot as plt
import nest
import numpy as np
import os
from pynestml.codegeneration.nest_code_generator_utils import NESTCodeGeneratorUtils
/home/charl/.local/lib/python3.11/site-packages/matplotlib/projections/__init__.py:63: UserWarning: Unable to import Axes3D. This may be due to multiple versions of Matplotlib being installed (e.g. as a system package and as a pip package). As a result, the 3D projection is not available.
warnings.warn("Unable to import Axes3D. This may be due to multiple versions of "
-- N E S T --
Copyright (C) 2004 The NEST Initiative
Version: 3.6.0-post0.dev0
Built: Mar 26 2024 08:52:51
This program is provided AS IS and comes with
NO WARRANTY. See the file LICENSE for details.
Problems or suggestions?
Visit https://www.nest-simulator.org
Type 'nest.help()' to find out more about NEST.
Paths
We assume here that we will generate code in a temporary directory /tmp/nestml-component. You can also create a unique temporary path using the Python tempfile module.
The Izhikevich model
A simple model for spiking neurons that nevertheless can exhibit a wide variety of dynamical behaviour, depending on its parameter values [1]. It is defined as follows:
\begin{align} \frac{dv}{dt} &= 0.04 v^2 + 5 v + 140 - u + I\\ \frac{du}{dt} &= a (b v - u) \end{align}
State update:
- :nbsphinx-math:`begin{align}
&text{if};; v geq V_{th}:\ &;;;; v text{ is set to } c\ &;;;; u text{ is incremented by } d\ & , \ &v text{ jumps on each spike arrival by the weight of the spike}
end{align}`
Example parameters for regular spiking (the meaning of these parameters is described in detail in the paper [1]; see also Task 2 below):
\begin{align} a&=0.02\\ b&=0.2\\ c&=-65~\text{mV}\\ d&=8 \end{align}
Task 1: Finish the model
In the file `izhikevich_task.nestml <https://raw.githubusercontent.com/nest/nestml/master/doc/tutorials/izhikevich/izhikevich_task.nestml>`__, only a subset of the parameters, state equations and update block is implemented.
Open the file in a text editor and finish the partially-completed model.
For reference, the solution is included as `izhikevich_solution.nestml <https://raw.githubusercontent.com/nest/nestml/master/doc/tutorials/izhikevich/izhikevich_solution.nestml>`__.
NESTML code generation
Assume that our NESTML input model is at izhikevich_solution.nestml. To generate code and build a dynamic library that can be loaded as a user module in NEST Simulator:
[2]:
# generate and build code
module_name, neuron_model_name = \
NESTCodeGeneratorUtils.generate_code_for("izhikevich_solution.nestml")
-- N E S T --
Copyright (C) 2004 The NEST Initiative
Version: 3.6.0-post0.dev0
Built: Mar 26 2024 08:52:51
This program is provided AS IS and comes with
NO WARRANTY. See the file LICENSE for details.
Problems or suggestions?
Visit https://www.nest-simulator.org
Type 'nest.help()' to find out more about NEST.
[12,izhikevich_tutorial_neuron_nestml, WARNING, [12:8;12:17]]: Variable 'a' has the same name as a physical unit!
[13,izhikevich_tutorial_neuron_nestml, WARNING, [15:8;15:17]]: Variable 'd' has the same name as a physical unit!
[17,izhikevich_tutorial_neuron_nestml, WARNING, [12:8;12:17]]: Variable 'a' has the same name as a physical unit!
[18,izhikevich_tutorial_neuron_nestml, WARNING, [15:8;15:17]]: Variable 'd' has the same name as a physical unit!
[23,izhikevich_tutorial_neuron_nestml, WARNING, [12:8;12:17]]: Variable 'a' has the same name as a physical unit!
[24,izhikevich_tutorial_neuron_nestml, WARNING, [15:8;15:17]]: Variable 'd' has the same name as a physical unit!
CMake Warning (dev) at CMakeLists.txt:93 (project):
cmake_minimum_required() should be called prior to this top-level project()
call. Please see the cmake-commands(7) manual for usage documentation of
both commands.
This warning is for project developers. Use -Wno-dev to suppress it.
-- The CXX compiler identification is GNU 12.3.0
-- Detecting CXX compiler ABI info
-- Detecting CXX compiler ABI info - done
-- Check for working CXX compiler: /usr/bin/c++ - skipped
-- Detecting CXX compile features
-- Detecting CXX compile features - done
-------------------------------------------------------
nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module Configuration Summary
-------------------------------------------------------
C++ compiler : /usr/bin/c++
Build static libs : OFF
C++ compiler flags :
NEST compiler flags : -std=c++17 -Wall -fopenmp -O2 -fdiagnostics-color=auto
NEST include dirs : -I/home/charl/julich/nest-simulator-install/include/nest -I/usr/include -I/usr/include -I/usr/include
NEST libraries flags : -L/home/charl/julich/nest-simulator-install/lib/nest -lnest -lsli /usr/lib/x86_64-linux-gnu/libltdl.so /usr/lib/x86_64-linux-gnu/libgsl.so /usr/lib/x86_64-linux-gnu/libgslcblas.so /usr/lib/gcc/x86_64-linux-gnu/12/libgomp.so /usr/lib/x86_64-linux-gnu/libpthread.a
-------------------------------------------------------
You can now build and install 'nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module' using
make
make install
The library file libnestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module.so will be installed to
/tmp/nestml_target_sii6g4x1
The module can be loaded into NEST using
(nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module) Install (in SLI)
nest.Install(nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module) (in PyNEST)
CMake Warning (dev) in CMakeLists.txt:
No cmake_minimum_required command is present. A line of code such as
cmake_minimum_required(VERSION 3.26)
should be added at the top of the file. The version specified may be lower
if you wish to support older CMake versions for this project. For more
information run "cmake --help-policy CMP0000".
This warning is for project developers. Use -Wno-dev to suppress it.
-- Configuring done (0.5s)
-- Generating done (0.0s)
-- Build files have been written to: /home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target
[ 33%] Building CXX object CMakeFiles/nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module_module.dir/nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module.o
[ 66%] Building CXX object CMakeFiles/nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module_module.dir/izhikevich_tutorial_neuron_nestml.o
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp: In member function ‘void izhikevich_tutorial_neuron_nestml::init_state_internal_()’:
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp:184:16: warning: unused variable ‘__resolution’ [-Wunused-variable]
184 | const double __resolution = nest::Time::get_resolution().get_ms(); // do not remove, this is necessary for the resolution() function
| ^~~~~~~~~~~~
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp: In member function ‘virtual void izhikevich_tutorial_neuron_nestml::update(const nest::Time&, long int, long int)’:
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp:318:24: warning: comparison of integer expressions of different signedness: ‘long int’ and ‘const size_t’ {aka ‘const long unsigned int’} [-Wsign-compare]
318 | for (long i = 0; i < NUM_SPIKE_RECEPTORS; ++i)
| ~~^~~~~~~~~~~~~~~~~~~~~
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp:313:10: warning: variable ‘get_t’ set but not used [-Wunused-but-set-variable]
313 | auto get_t = [origin, lag](){ return nest::Time( nest::Time::step( origin.get_steps() + lag + 1) ).get_ms(); };
| ^~~~~
/home/charl/julich/nestml-fork-integrate_specific_odes/nestml/doc/tutorials/izhikevich/target/izhikevich_tutorial_neuron_nestml.cpp:303:16: warning: unused variable ‘__resolution’ [-Wunused-variable]
303 | const double __resolution = nest::Time::get_resolution().get_ms(); // do not remove, this is necessary for the resolution() function
| ^~~~~~~~~~~~
[100%] Linking CXX shared module nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module.so
[100%] Built target nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module_module
[100%] Built target nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module_module
Install the project...
-- Install configuration: ""
-- Installing: /tmp/nestml_target_sii6g4x1/nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module.so
Check the generated log output for any potential error messages or warnings.
Instantiate model in NEST Simulator and run
The generated extension module can be loaded using nest.Install().
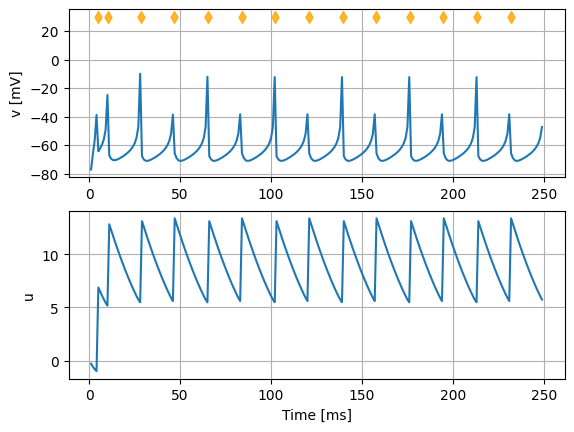
Using the PyNEST API, the model can be instantiated and simulated in NEST. The following code will create one instance of the neuron model (nest.Create("izhikevich_tutorial")), inject a constant current and run the simulation for 250 ms.
[3]:
nest.set_verbosity("M_ALL")
nest.ResetKernel()
# load dynamic library (NEST extension module) into NEST kernel
nest.Install(module_name)
neuron = nest.Create(neuron_model_name)
voltmeter = nest.Create("voltmeter")
voltmeter.set({"record_from": ["v", "u"]})
nest.Connect(voltmeter, neuron)
cgs = nest.Create('dc_generator')
cgs.set({"amplitude": 25.})
nest.Connect(cgs, neuron)
sr = nest.Create("spike_recorder")
nest.Connect(neuron, sr)
nest.Simulate(250.)
spike_times = nest.GetStatus(sr, keys='events')[0]['times']
fig, ax = plt.subplots(nrows=2)
ax[0].plot(voltmeter.get("events")["times"], voltmeter.get("events")["v"])
ax[1].plot(voltmeter.get("events")["times"], voltmeter.get("events")["u"])
ax[0].scatter(spike_times, 30 * np.ones_like(spike_times), marker="d", c="orange", alpha=.8, zorder=99)
for _ax in ax:
_ax.grid(True)
ax[0].set_ylabel("v [mV]")
ax[1].set_ylabel("u")
ax[-1].set_xlabel("Time [ms]")
fig.show()
Apr 19 10:44:47 Install [Info]:
loaded module nestml_004fd3f5ec9a44b9a28667ae47e2bc9e_module
Apr 19 10:44:47 NodeManager::prepare_nodes [Info]:
Preparing 4 nodes for simulation.
Apr 19 10:44:47 SimulationManager::start_updating_ [Info]:
Number of local nodes: 4
Simulation time (ms): 250
Number of OpenMP threads: 1
Not using MPI
Apr 19 10:44:47 SimulationManager::run [Info]:
Simulation finished.
/tmp/ipykernel_329471/1245292433.py:33: UserWarning:FigureCanvasAgg is non-interactive, and thus cannot be shown
Task 2: Parameter space exploration
Perform a parameter space exploration to reproduce the bottom eight panels from [1], figure 2.
References
[1] Eugene M. Izhikevich, “Simple Model of Spiking Neurons”, IEEE Transactions on Neural Networks, Vol. 14, No. 6, November 2003
Copyright
This file is part of NEST.
Copyright (C) 2004 The NEST Initiative
NEST is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 2 of the License, or (at your option) any later version.
NEST is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with NEST. If not, see http://www.gnu.org/licenses/.